Genes Responding to Oxidative Stress in Wild Type and sty1 Deletion S.Pombe
Abstract
Sty1 is a mitogen-activated protein kinase (MAPK) from the yeast Schizosaccharomyces pombe which is homologous to the human MAPK p38. It is activated by several stress responses including oxidative stress. In this study the transcriptomes of a wild type S. pombe strain and a sty1 delete (Δsty1) strain were studied using microarray technology and were compared. By looking at differences in up- and downregulation between the two strains when subjected to oxidative stress in the presence of hydrogen peroxide, genes that are regulated by sty1 were found. Their functions and any human orthologs were found by using the online database, PomBase, and the human orthologs were researched to find links to the p38 MAPK pathway. One of these genes, SPBC1685.05, a serine protease, was found to be homologous to the HtrA family of proteases. It was found that HTRA2 is activated by p38 via PINK1 and CDK5 and is involved in cleavage of misfolded proteins as it can selectively bind to them using a PDZ domain. It was also noted that mutations in the HTRA2 gene can be linked to Parkinson’s disease, as can mutations in the closely related gene for PINK1, another kinase known to regulate HTRA2.
Introduction
Schizosaccharomyces pombe is a species of yeast commonly used as a model organism in molecular and cellular biology as it is a relatively simple, single-celled eukaryote (reviewed by Yanagida, 2002). Because of this, a lot of its genes are homologous to human genes so studying these genes through overexpression or deletion can often give clues as to the function of the homologous human genes (Choi et al., 2004). In this study sty1, also known as spc1, is studied by comparing gene expression in wild type S. pombe cells with a strain in which sty1 has been deleted in order to find genes that are regulated by sty1.
It is important that all organisms, including S. pombe, are able to respond efficiently to environmental stresses such as oxidative or osmotic stress in order to survive harsh environmental conditions. Oxidative stress is caused by reactive oxygen species (ROS), such as peroxides, in excess of the level that a cell or organism can ordinarily detoxify which results in damage (Valko et al., 2005). The sty1 pathway is activated, as discussed above, with peroxides being sensed by the histidine kinases Mak2 and 3 which then eventually activate sty1 via the phosphorylation and MAP kinase cascades.
The mitogen-activated protein kinase (MAPK) cascade is important in all eukaryotic cells in the response to environmental stresses, such as oxidative stress. MAP kinases were first sequenced in 1989 by Courchesne et al. (1989). In the early 1990s, from the study of different species, it was discovered that there are multiple MAP kinases. MAP kinases are activated by a further two protein kinases acting in sequence. In all known cases the MAP kinase (MKK or MEK) phosphorylates serine/threonine and tyrosine residues (Ahn et al., 1991). MEKs are themselves phosphorylated by MEK kinases (MEKKs) at two serine or threonine residues (Zheng and Guan, 1994).
Sty1 is a gene that codes for a MAP Kinase (Sty1) activated by the MEK Wis1 (Warbrick and Fantes, 1991) via phosphorylation of threonine 171 and tyrosine 173 (Shiozaki and Russell, 1995). Wis1 is itself phosphorylated by Wis4 (Samejima et al., 1997) and Win1 (Shieh et al., 1998a). However, Shieh et al. (1998b) showed that when a mutant wis1 is overexpressed sty1 is still activated by stress although the mechanism is unknown. Wis4 and win1 are activated by a response regulator, Mcs4. This is controlled by the phosphotransmitter, Spy1, itself activated by the histidine kinases Mak2 and 3, which sense peroxide. The phosphorylation cascade is known to be activated by several stress responses, as discovered by Degols, Shiozaki and Russell (1996), although it was originally thought to be activated only by osmotic stress like its close relation, Saccharomyces cerevisiae, hog1. It is now thought that not only does Mcs4 receive stress signals from the Mak2/3-Spy1 cascade, but it stabilises the heteromer formed by Wis4 and Win1, promoting interaction with Wis1 (Morigasaki et al., 2013).
MAP kinases activate transcription factors which regulate the transcription of genes. Sty1, which accumulates in the nucleus once phosphorylated, regulates the transcription of a large number of genes, mostly via the transcription factor Atf1 which forms a heterodimer with Pcr1 (Reiter et al., 2008). Although sty1 phosphorylates Atf1, Reiter et al. (2008) showed that this is not critical for transcriptional activation; rather, sty1 is recruited to promoters of Atf1-dependant genes.
The sty1 pathway is homologous to the human p38 MAP kinase pathway. There are four P38 MAP kinases (p38α,β,γ and δ), or MAPK11(β),12(γ),13(δ) and 14(α) (also known as p38) which are usually phosphorylated by MEK3 or MEK6 but have also been shown to be phosphorylated by MEK4 as part of a complex network (figure 1).
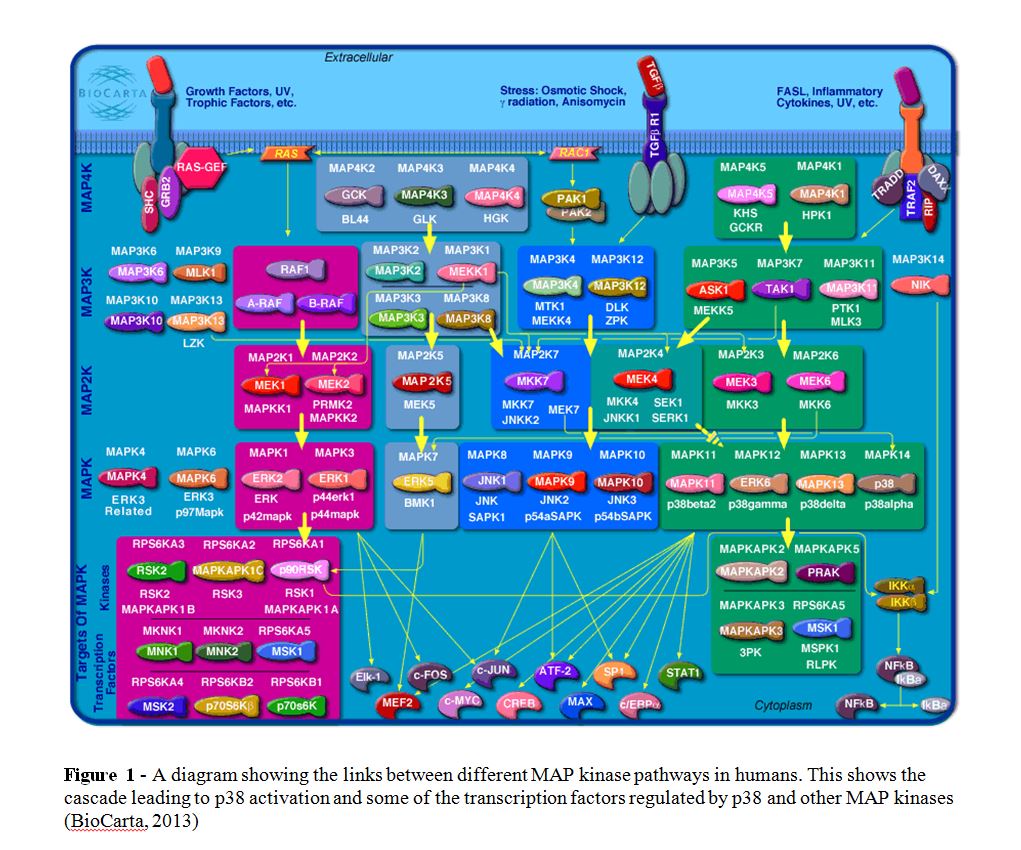
Genes involved in or regulated by the cascade can often be related to disease. Mutations in the gene generally cause an incorrect amino acid in the protein product sequence and hence a misfolding of the protein, which is likely to disrupt its function. These genes are often involved in the stress response. Diseases develop due to an inability to cope with the stress presented; for example, removing excess radicals or removing other misfolded proteins that can then form amyloid plaques (reviewed by Herczenik and Gebbink, 2008).
In this investigation, genes in dataset 3 from the transcriptomes of wild type and Δsty1 S. pombe were compared under conditions of oxidative stress. From this it can be determined which genes are regulated by the sty1 MAP kinase pathway by comparing data from the wild type cells and the Δsty1 cells where the pathway has been inactivated. Online databases can then be used to find more information about these genes. From this information, a link to MAP kinase pathways can be inferred by looking at orthologs from other species. This report aims to find these genes and use the available information from online resources to either find a direct link or propose a link to the sty1 pathway. Links between human orthologs and specific diseases will also be investigated.
Methods
In this experiment, microarray analysis was carried out on the total RNA from both wild type and Δsty1 S.pombe yeast cells. Both strains were subjected to oxidative stress (2mM H2O2) and samples collected at the start and after 15 and 60 minutes. The total RNA was isolated and labelled and underwent microarray hybridisation, as described by Lyne et al. (2003). Microarray data for each strain was obtained in duplicate and the results normalised to the values obtained at time 0. For each gene, up- or down-regulation was assessed by averaging the two values for each gene and calculating the ratio of fluorescence at 15 and 60 minutes to that at 0 minutes. From this the fold change was calculated as log to the base 2 of the ratios. It was decided that significant up- or down regulation would be when the fold change after 60 minutes was greater than 2 or lower than -2 in order to limit the number of genes researched. From this, a large number of genes were found to be significantly regulated by sty1 and more information about these genes, including the function of their protein products and the presence of human orthologs, was found using the PomBase online database.
Results
A large number of genes were identified that were significantly up-regulated under oxidative stress conditions in the wild type cells but not in the Δsty1 cells, suggesting that these genes are activated by sty1. The most significant of these was SPBC24C6.09c, thought to be a phosphoketolase, which does not appear to have a human ortholog. The fold changes of the most significant ones that do have human orthologs are summarised in Table 1.
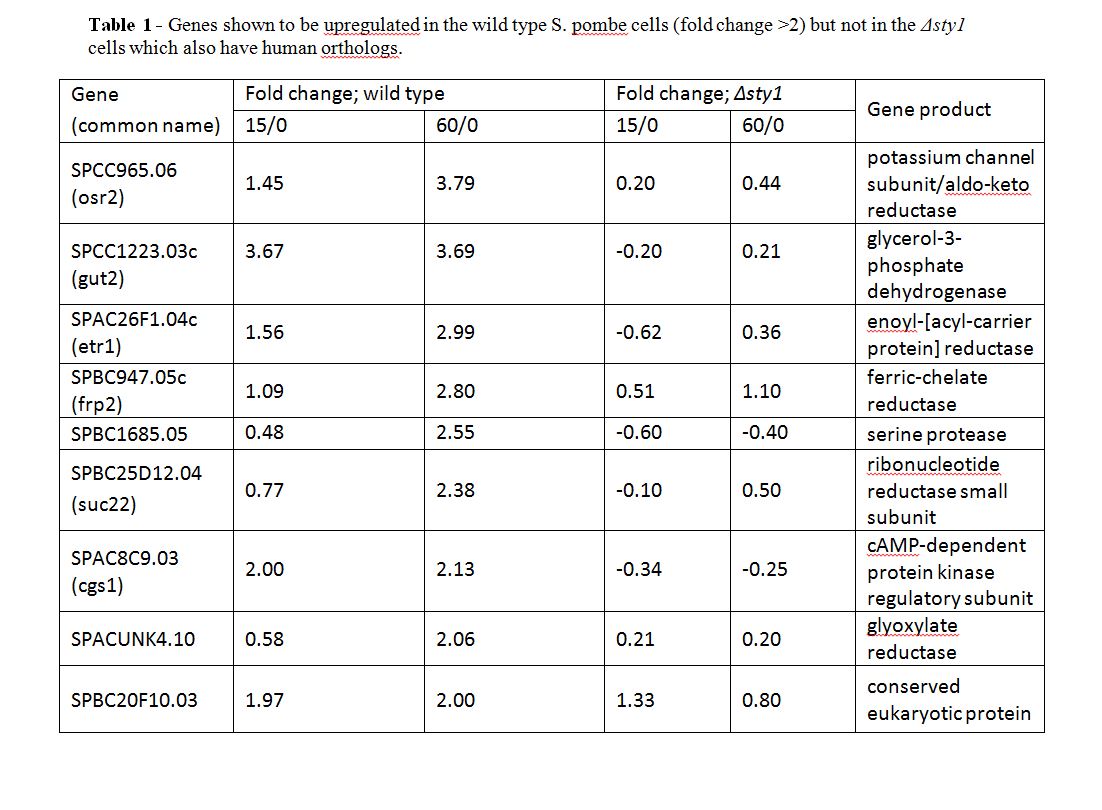
These fold changes were obtained from the ratio of the microarray data after 15 or 60 minutes of oxidative stress compared to no oxidative stress. The data for SPBC1685.05 is exemplified below (Figure 2) as this gene will be further discussed later in this report.
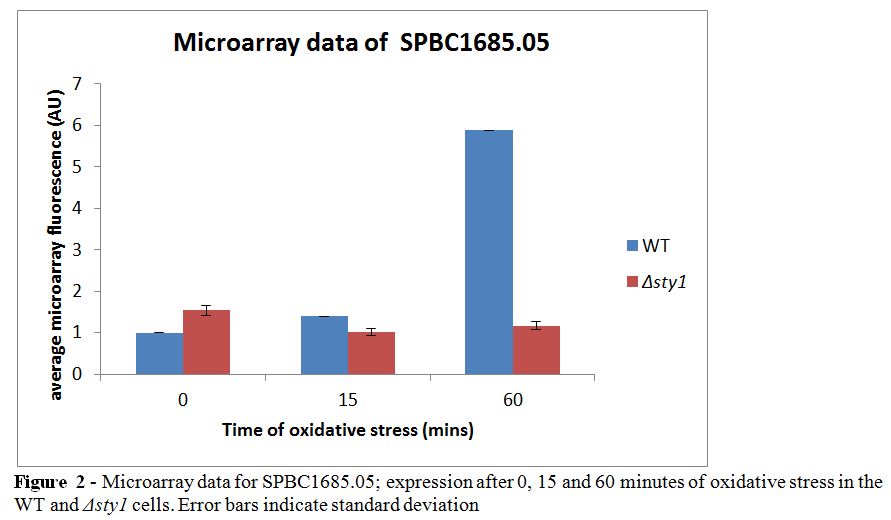
Five genes were found to be significantly down-regulated under oxidative stress conditions in the wild type but not the Δsty1 cells, all of which have a human ortholog (see Table 2) including SPBC17D1.06, also known as dbp3, a predicted ATP-dependant RNA helicase which has several functions in RNA processing.
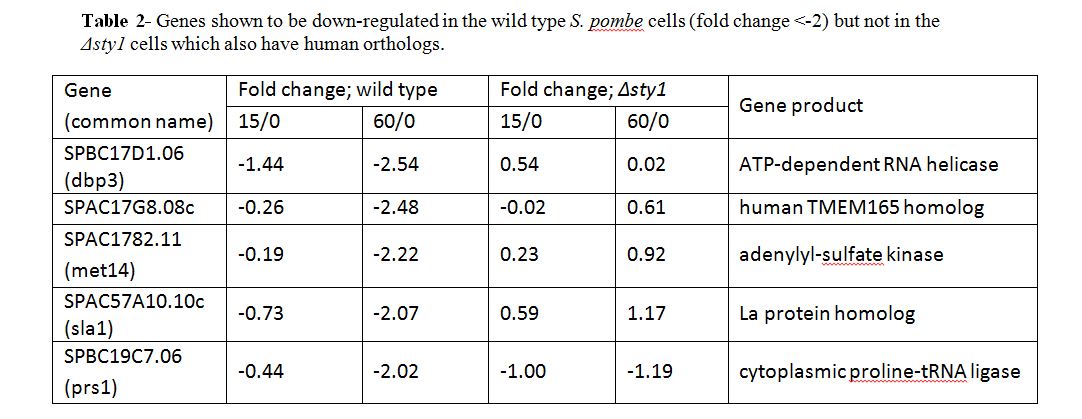
There were also some genes that were not affected by oxidative stress in the wild type but were upregulated in the Δsty1 cells, suggesting that these genes are repressed by sty1 but not necessarily under stress conditions. However, none of these have a human ortholog. Eight genes were down-regulated in the Δsty1 cells under stress but were not affected by stress in the wild type cells. Seven of these have a human ortholog (see Table 3), including SPAC20G8.06, also known as not1, which is predicted to code for a subunit of the CCR4-Not complex. It is implied that they are up-regulated by sty1 but not necessarily under stress.
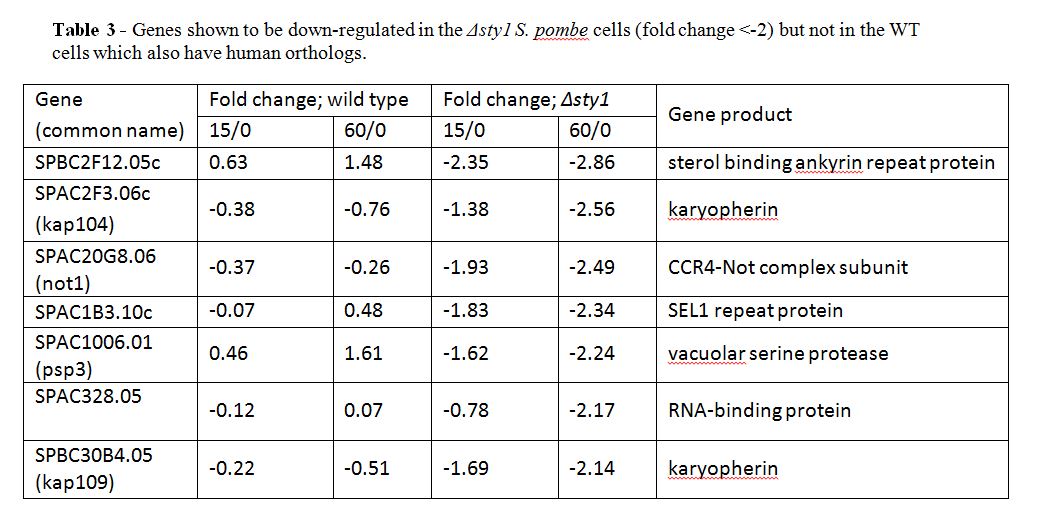
As well as the genes outlined in Tables 1−3 other genes were shown to be regulated by sty1. However, they were either not as significantly up- or down-regulated or they did not have human orthologs. There were also some genes that were up- or down-regulated in both the wild type and the Δsty1 cells and hence were shown not to be regulated by sty1. They were also disregarded for the purposes of this study. However, all genes shown to be up- or down-regulated are summarised in the appendices.
Discussion
There were some obvious trends in the expression pattern of S. pombe genes when subjected to oxidative stress. For example, there appear to be a lot of oxidoreductases upregulated; however this is unsurprising, given that they act to detoxify free oxygen radicals which are the cause of oxidative stress (Kawasaki et al., 2009). Genes involved in carbohydrate metabolism also appear to be up-regulated, which again is unsurprising as carbohydrate metabolism in the form of glycolysis and the pentose phosphate pathway is a good source of NADH and NADPH. NADPH can aid the reduction of oxidised glutathione (GSSG) to its reduced form (GSH) via NADPH-dependant glutathione reductase. GSH reacts with H2O2 and has been shown to play an important part in adaption to oxidative stress (Salvemini et al., 1999).
Of the analysed genes, few that were significantly regulated by sty1 also had human orthologs. Of the orthologous genes even fewer appeared to be directly affected by the human ortholog of sty1, p38. While it is only possible to discuss one of these genes in detail in this report, all genes that are up- or down-regulated to any great extent in either the wild type, Δsty1 or both cell strains, are listed in the appendices.
SPBC1685.05 is a gene that was up-regulated in the wild type cells but not the Δsty1 cells. It codes for a serine proteinase which is homologous to the HTRA family of proteases in humans, HTRA1-4 (PomBase, 2013). Little appears to be known about the protein product of SPBC1685.05 (herein SPBC1685.05); however, an important feature is its PDZ domain, which is a highly conserved protein-binding domain also found in the HTRA proteins (Clausen et al., 2002).
It has been shown that high-temperature regulated A2 (HTRA2), a mitochondrial protease that is part of the HTRA family, also known as Omi, is regulated by the MAP Kinase p38, which is homologous to sty1 (Faccio et al., 2000). In times of oxidative stress the presence of reactive oxygen species (ROS) can damage several cellular components resulting in an increase in misfolded proteins. One of the probable functions of HTRA2 is to cleave and hence destroy these proteins (Clausen et al., 2011). Therefore it is inferred that HTRA2 should be up-regulated while subjected to oxidative stress.
HTRA2 is activated by phosphorylation by PTEN-induced putative kinase 1 (PINK1), or cyclin-dependent kinase 5 (CDK5), themselves promoted by p38 (Plun-Favreau et al., 2007; Fitzgerald et al., 2012). While a definite ortholog of PINK1 does not appear to have been found in S. pombe, it is potentially orthologous to ksp1. CDK5 is homologous to the S. pombe pef1, although neither gene seems to be linked to sty1 (PomBase, 2013).
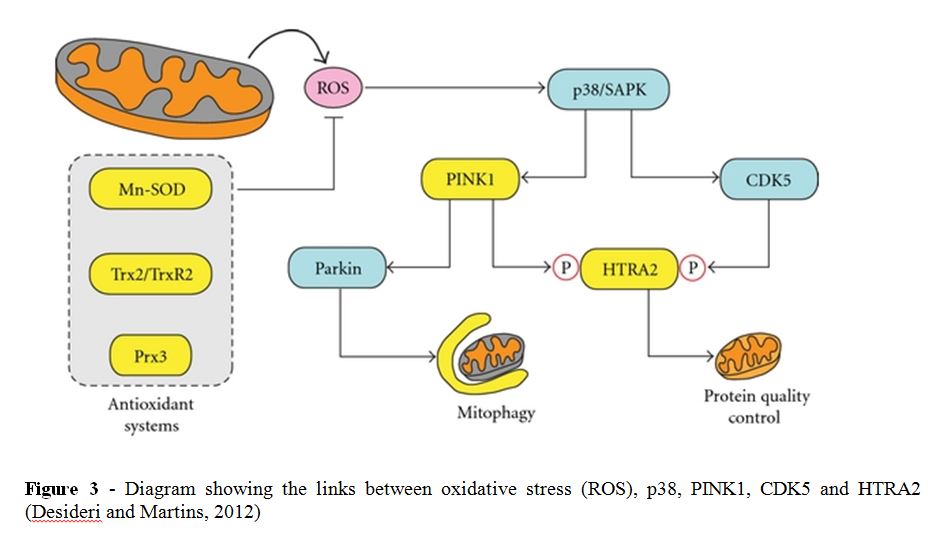
Given that SPBC1685.05 is homologous to HTRA2, which presumably has the same or a similar function, it can be understood why it is upregulated in the wild type S. pombe cells when subjected to oxidative stress. As HTRA2 is regulated by p38, which is homologous to sty1, it can be inferred that SPBC1685.05 is regulated by the kinase sty1. Therefore it makes sense that in the Δsty1 cells, which lack this pathway, SPBC1685.05 is no longer up-regulated. However, the link between HTRA2 and p38 is via phosphorylation of the gene product rather than by up-regulation of the gene itself. Thus there could be another mechanism of up-regulation that does not appear to have been explored.
As expected due to its homology with SPBC1685.05, HTRA2 also has a PDZ domain. This is important as it acts as a regulator of the protein’s proteolytic activity. The PDZ regulates proteolysis by only allowing it when bound to specific PDZ-binding proteins or internal hydrophobic stretches of misfolded proteins. This is how the protein knows to selectively cleave misfolded proteins and not just any available protein (Zhang et al., 2007). It is possible that this domain in SPBC1685.05 has the same function.
It has been shown that HTRA2 is linked to Parkinson’s disease in certain sporadic cases with certain loss-of-function mutations causing the disease. The importance of the PDZ domain is emphasised here as certain mutations cause the condition (Strauss et al., 2005). This is logical given that a mutation here would stop binding to and hence cleavage of misfolded proteins. They would build up, causing the condition.
To determine whether SPBC1685.05 has the same function as HTRA2, this could be tested using recombinant yeast where SPBC1685.05 is knocked out. HTRA2 is inserted into the genome in its place and the phenotype compared to the WT. What is interesting is that the other HTRA proteins do not appear to be regulated by p38. If studies testing their relationship have not been conducted, this could be a consideration for the future. It is also interesting that SPAC23G3.12c, another S.pombe gene that is apparently homologous to the HTRA family of proteases, is not significantly regulated by sty1, so it is possible that this gene’s product is more homologous to the other members of the family if it is not regulated by p38.
In conclusion, many genes were found to be regulated by sty1 with some of them having human orthologs that could have been shown to be regulated by p38 which is homologous to sty1. One of these, SPBC1685.05, was discussed in detail including its function, human ortholog and the diseases associated with it.
References
Ahn, N.G., Seger, R., Bratlien, R.L., Diltz, C.D., Tonks, N.K. and Krebs, E.G., 1991, Multiple components in an epidermal growth-factor stimulated protein kinase cascade. In vitro activation of a myelin basic protein/microtubule-associated protein 2 kinase, Journal of Biological Chemistry, 266(7), 4220-4227
BioCarta, 2013, MAPKinase signalling pathway, [online] available from http://www.biocarta.com/pathfiles/h_mapkpathway.asp, accessed 12/11/13
Choi, W.S., Lin, Y.C. and Gralla, J.D., 2004, The Schizosaccharomyces pombe Open Promoter Bubble: Mammalian-like Arrangement and Properties, Journal of Molecular Biology, 340(5), 981-989
Clausen, T., Southan, C. and Ehrmann, M., 2002, The HtrA family of proteases: implications for protein composition and cell fate, Molecular Cell, 10(3), 443-455
Clausen, T., Kaiser, M., Huber, R. and Ehrmann, M., 2011, HTRA proteases: regulated proteolysis in protein quality control, Nature Reviews Molecular Cell Biology, 12(3), 152–162
Courchesne, W.E., Kunisawa, R. and Thorner, J., 1989, A putative protein kinase overcomes pheromone-induced arrest of cell cycling in S. cerevisiae, Cell, 58(6), 1107-1119
Degols, G., Shiozaki, K. and Russell, P., 1996, Activation and Regulation of the Spc1 Stress-Activated Protein Kinase in Schizosaccharomyces pombe, Molecular and Cellular Biology, 16(6) 2870-2877
Desideri, E. and Martins, M., 2012, Mitochondrial Stress Signalling: HTRA2 and Parkinson’s disease, International Journal of Cell Biology, 2012, Article ID 607929
Faccio, L., Fusco, C., Chen, A., Martinotti, S., Bonventre, J.V. and Zervos, A.S., 2000, Characterization of a Novel Human Serine Protease That Has Extensive Homology to Bacterial Heat Shock Endoprotease HtrA and Is Regulated by Kidney Ischemia, Journal of Biological Chemistry, 275(4), 2581-2588
Fitzgerald, J.C., Camprubi, M.D., Dunn, L., Wu, H-C., Ip, N.Y., Kruger, R., Martins, L.M., Wood, N.W. and Plun-Favreau, H., 2012, Phosphorylation of HtrA2by cyclin-dependant kinase 5 is important for mitochondrial function, Cell Death and Differentiation, 19(2012), 257-266
Herczenik, E and Gebbink, F.B.G., 2008, Molecular and cellular aspects of protein misfolding and disease, The FASEB Journal, 22(7), 2115-2133
Kawasaki, S., Sakai, Y., Takahasi, T., Suzuki, I. and Niimura, Y., 2009, O2 and Reactive Oxygen Species Detoxification Complex, Composed of O2-Responsive NADH:Rubredoxin Oxidoreductase-Flavoprotein A2-Desulfoferrodoxin Operon Enzymes, Rubperoxin, and Rubredoxin, in Clostridium acetobutylicum, Applied and Environmental Microbiology, 75(4), 1021-1029
Lyne, R., Burns, G., Mata, J., Penkett, C.J., Rustici, G., Chen, D., Langford, C., Vetrie, D. and Bähler, J., 2003, Whole-genome microarrays of fission yeast: characteristics, accuracy, reproducibility, and processing of array data, BMC Genomics, 4(1), 27-42
Morigasaki, S., Ikner, A., Tatebe, H. and Shiozaki, K., 2013, Response regulator-mediated MAPKKK heteromer promotes stress signaling to the Spc1 MAPK in fission yeast, Molecular Biology of the Cell, 24(7), 1083-1092
Plun-Favreau, H., Klupsch, K., Moisoi, N., Gandhi, S., Kjaer, S., Frith, D., Harvey, K., Deas, E., Harvey, R.J., McDonald, N., Wood, N.W., Martins, L.M., Downward, J., 2007, The mitochondrial protease HtrA2 is regulated by Parkinson’s disease-associated kinase PINK1, Nature Cell Biology, 9(11), 1243-1252
PomBase, 2013, PomBase, [online] available from http://www.pombase.org/ accessed 12/11/13
Reiter, W., Watt, S., Dawson, K., Lawrence, C.L., Bähler, J., Jones, N. and Wilkinson, C.R.M., 2008, Fission Yeast MAP Kinase Sty1 Is Recruited to Stress-induced Genes, Journal of Biological Chemistry, 283(15), 9945-9956
Salvemini, F., Franze, A., Iervolino, A., Filosa, S., Salzano, S. and Ursini, M.V., 1999, Enhanced Glutathione Levels and Oxidoresistance Mediated by Increased Glucose-6-phosphate Dehydrogenase Expression, Journal of Biological Chemistry, 274(5), 2750-2757
Samejima, I., Mackie, S. and Fantes, P.A., 1997, Multiple modes of activation of the stress-responsive MAP kinase pathway in fission yeast, EMBO Journal, 16(20), 6162-6170
Shieh, J-C., Wilkinson, M.G. and Millar, J.B.A., 1998a, The Win1 Mitotic Regulator Is a Component of the Fission Yeast Stress-activated Sty1 MAPK Pathway, Molecular Biology of the Cell, 9(2), 311-322
Shieh, J-C., Marin, H. and Millar, J.B.A., 1998b, Evidence for a novel MAPKKK-independent pathway controlling the stress activated Sty1/Spc1 MAP kinase in fission yeast, Journal of Cell Science, 111(1998), 2799-2807
Shiozaki, K. and Russell, P., 1995, Cell-cycle control linked to the extracellular environment by MAP kinase pathway in fission yeast, Nature (London), 378, 739-743
Strauss, K.M., Martins, L.M., Plun-Favreau, H., Marx, F.P., Kautzmann, S., Berg, D., Gasser, T., Wszolek, Z., Muller, T., Bornemann, A., Wolburg, H., Downward, J., Riess, O., Schulz, J.B. and Kruger, R., 2005, Loss of function mutations in the geneencoding Omi/HtrA2 in Parkinson’s disease, Human Molecular Genetics, 14(15), 2099-2111
Valko, M., Morris, H. and Cronin, M.T.D., 2005, Metals, Toxicity and Oxidative Stress, Current Medicinal Chemistry, 12(10), 1161-1208
Warbrick, E. and Fantes, P.A., 1991, The wis1 protein is a dosage-dependant regulator of mitosis in Schizosacchromyces pombe, EMBO Journal, 10(13), 4291-4299
Yanagida, M., 2002, The model unicellular eukaryote, Schizosaccharomyces pombe, Genome Biology, 3(3), 1-4
Zhang, Y., Appleton, B.A., Wu, P., Wiesmann, C. and Sidhu, S.S., 2007, Structural and Functional Analysis of the ligand specificity of the HtrA2/Omi PDZ domain, Protein Science, 16(8), 1738-1750
Zheng, C-F. and Guan, K., 1994, Activation of MEK family kinases requires phosphorylation of two conserved Ser/Thr residues, EMBO Journal, 13(5) 1123-1131